Science
Scientific vision
Antibiotics have transformed modern medicine, yet many bacterial infections remain difficult to cure. In chronic respiratory pathogens frequently persist and cause treatment failures despite therapies being considered active in standard laboratory tests.
Our research aims to redefine how antibiotic responses are measured and understood. By combining new experimental and computational technologies with clinical research, we study how individual bacteria respond to treatment and how these responses shape treatment outcomes. Ultimately, our goal is to use these insights to design more effective antibiotic therapies and enable personalised strategies for difficult-to-treat infections.
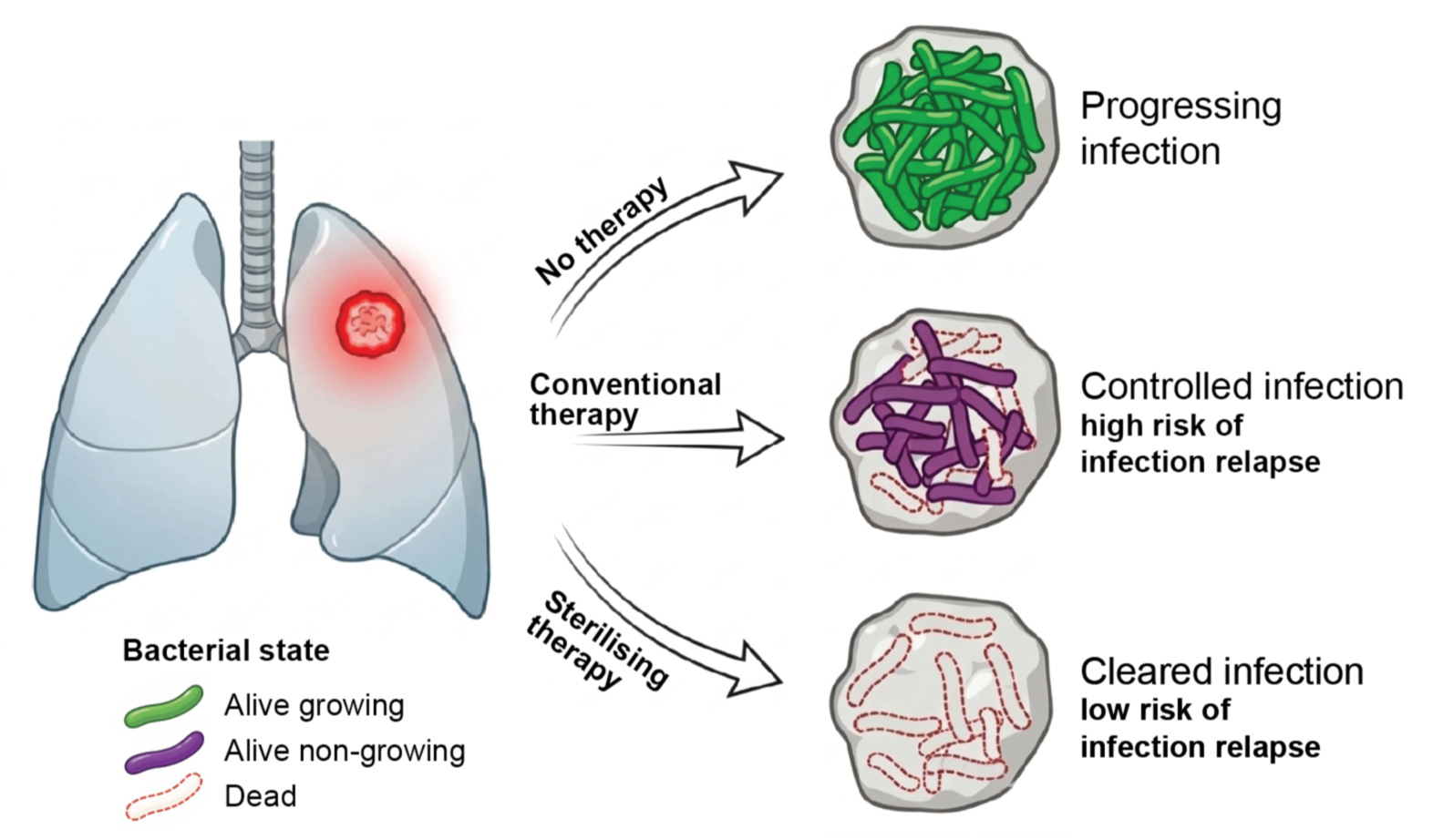
Respiratory infections
Pulmonary infections are among the most common infectious diseases worldwide and remain a major cause of morbidity and mortality. They frequently complicate hospital stays and the treatment of chronic diseases, for example in patients with bronchiectasis, COPD or immunosuppression. In addition, pulmonary infections account for the largest proportion of antibiotic resistance-related deaths.
Tuberculosis remains another major global challenge, causing more than ten million infections and over one million deaths each year. Despite this enormous disease burden, treatment of tuberculosis still relies on prolonged multidrug regimens.
While science and medicine have advanced dramatically in many fields, treatment strategies for pulmonary infections have hardly changed over the past decades.
The biology of bacterial survival
A key reason why many bacterial infections are difficult to cure is the ability of bacteria to survive antibiotic treatment. Even when exposed to drugs that are considered active (that means that drugs are able to inhibit bacterial growth) bacteria are often poorly killed. As a result, some bacteria are able to survive treatment and to subsequently resume growth once the antibiotic is removed, leading to prolonged infections, treatment failure, and relapse.
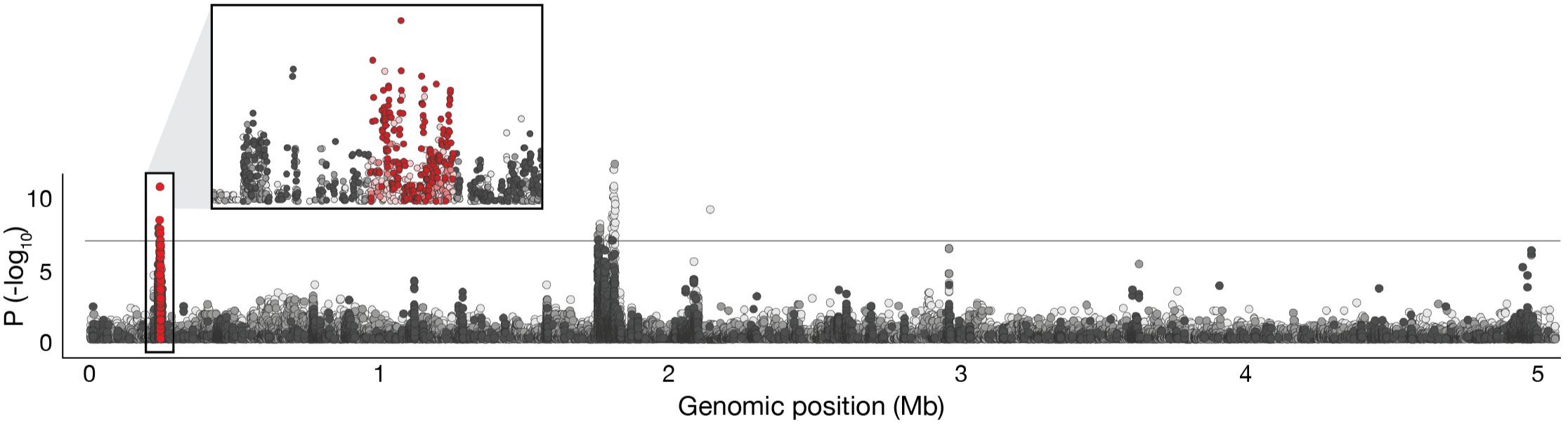
Understanding why bacteria survive treatment is therefore a central question in infection biology. In our lab, we aim to uncover the biological mechanisms that allow bacteria to withstand antibiotic exposure. By studying bacterial responses to antibiotics at high temporal and single-cell resolution and at large scale, we aim to identify the physiological states, genetic pathways, and environmental factors that shape bacterial survival during drug treatment. Our goal is to understand the processes that determine whether antibiotic treatment eradicates an infection or allows bacterial persistence.
Rational design of antibiotic treatments
Antibiotic discovery and development is expensive and prone to failure. The principles that guide the design of antibiotic therapy remain surprisingly limited. In many cases, drug combinations and treatment durations were established empirically, and the biological factors that determine treatment success are still poorly understood.
In our research, we use a rational framework to establish antibiotic treatments. We have shown that bacterial survival under nutrient-limited conditions is closely linked to treatment outcomes in mouse models and clinical trials. We therefore focus on identifying treatment targets and drug combinations with strong sterilising capacity, even when bacteria are metabolically inactive.
Our goal is to use these insights to improve treatments for non-tuberculous mycobacterial infections, shorten treatments for tuberculosis, and develop treatments that successfully eradicate chronic Pseudomonas aeruginosa infections, to achieve durable and long-lasting cures.
Personalised infection therapy
For decades antibiotic therapy has been guided by conventional antibiotic susceptibility testing, which measures bacterial growth inhibition under standardised laboratory conditions. However, these tests often fail to capture the complex responses of bacteria during antibiotic exposure and largely ignore other mechanisms that contribute to treatment failure, such as antibiotic tolerance and persistence.
In our work, we have shown that drug tolerance profiles vary substantially between clinical isolates and are linked to individual treatment outcomes in Mycobacterium abscessus pulmonary disease. This opens the possibility of tailoring therapies not only based on resistance, but also on tolerance patterns.
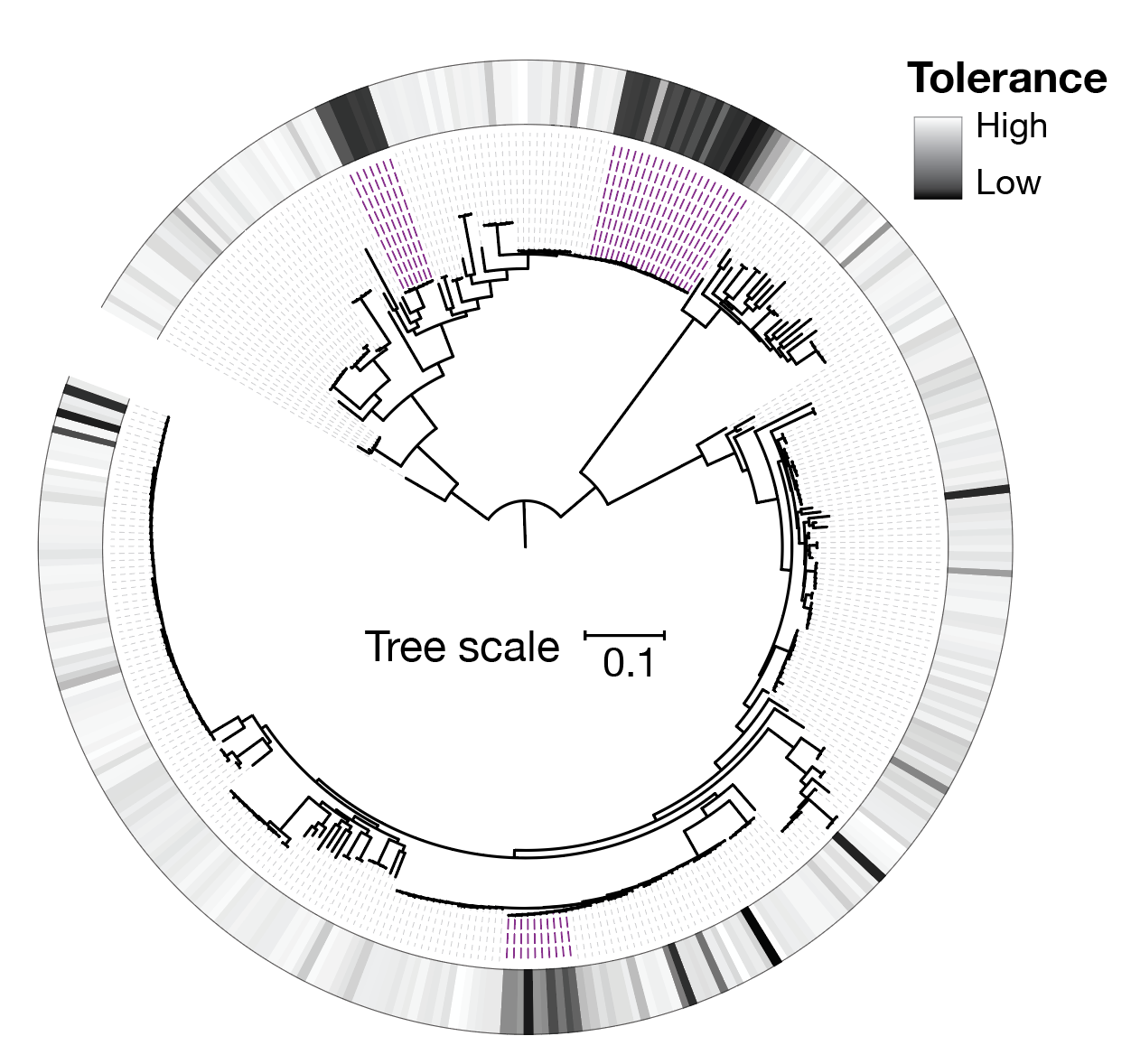
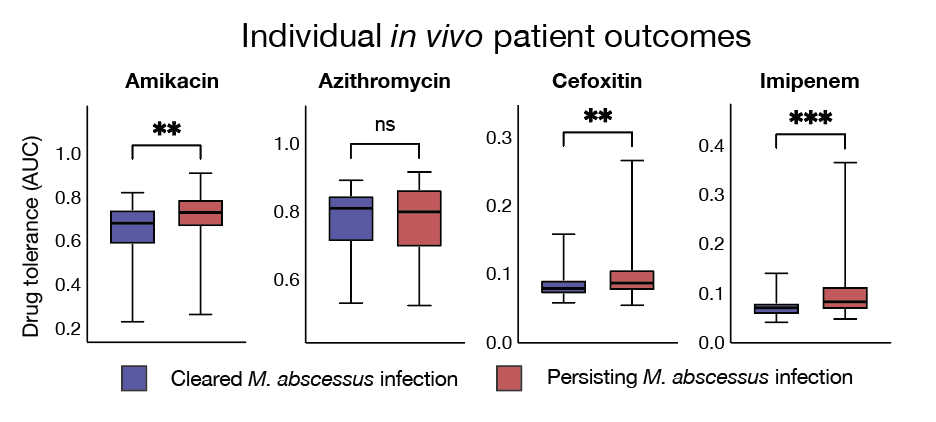
Our goal is to translate these insights into personalised therapy, where antibiotic treatments are tailored to the biological properties of individual infections and designed to maximise bacterial killing while preventing infection persistence.
Technologies
Knowledge and medical advances are often driven by technological innovation. In our lab, we develop new experimental methods and combine them with other cutting-edge technologies, ranging from CRISPR-based gene editing to genomic sequencing, proteomics, and advanced computational approaches. By embedding these technologies in clinical studies, we aim to develop new strategies to address some of the most challenging problems in infection treatment.
Antimicrobial single-cell testing. To accurately study antibiotic effects and bacterial escape strategies, we developed a high-content live-cell imaging approach that immobilises single bacteria at the bottom of multi-well plates. This strategy allows us to assess thousands of experimental conditions, track millions of bacteria over several days, and measure bacterial viability in real time. Importantly, these measurements (across different antibiotics and mycobacterial isolates) are linked to treatment outcomes in human infections.
AI-based image analysis. Our experimental platforms generate vast amounts of single-cell time-lapse data. To analyse these datasets, we develop computational approaches that allow us to extract biological information from millions of individual bacterial trajectories. Using supervised and unsupervised machine learning, we identify patterns of bacterial behaviour that would otherwise remain hidden.
Functional genomics. To understand the genetic basis underlying distinct bacterial behaviours, such as antibiotic survival, we use genome-wide functional genomics approaches. These include CRISPR-based perturbation libraries and phenogenomic analysis, an approach we developed together with Andres Floto’s team. Using these strategies, we systematically identify genes and pathways that shape bacterial physiology and antibiotic responses.
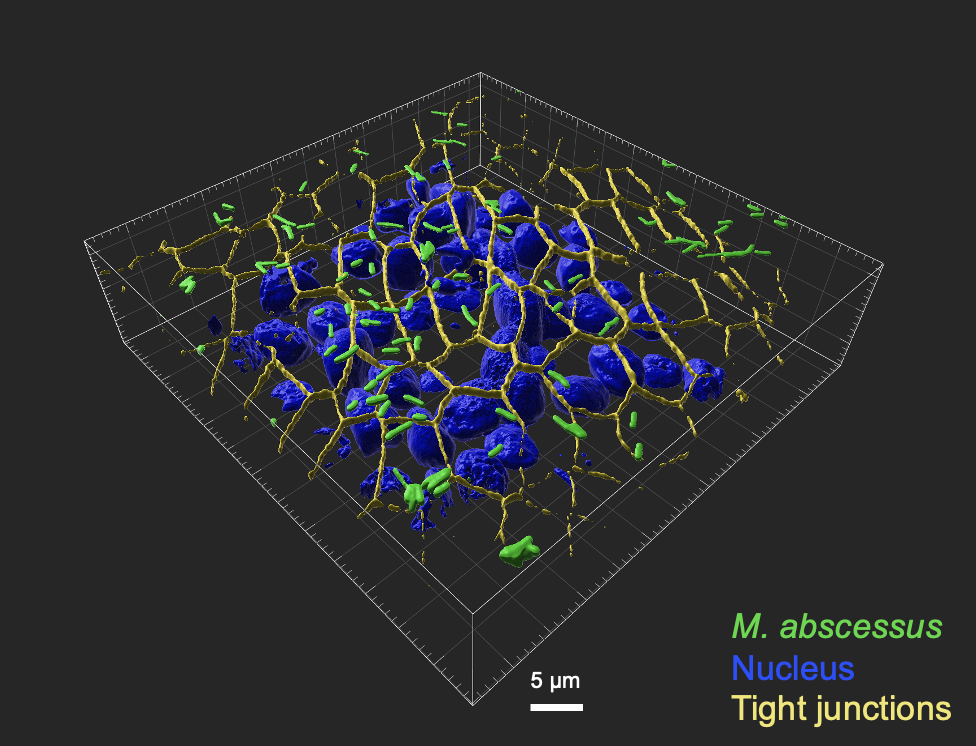
Translational infection models. To connect bacterial measurements with infections in patients, we study bacterial infections in a human-derived airway models. These air-liquid interface systems recapitulate the complex structure and cellular composition of the human airway epithelium and allow us to investigate host-pathogen interactions under physiologically relevant conditions. In collaboration with Urs Jenal’s team, we use these models to understand how bacteria behave in the lung environment and how antibiotic treatments perform in conditions that closely resembles human infection.
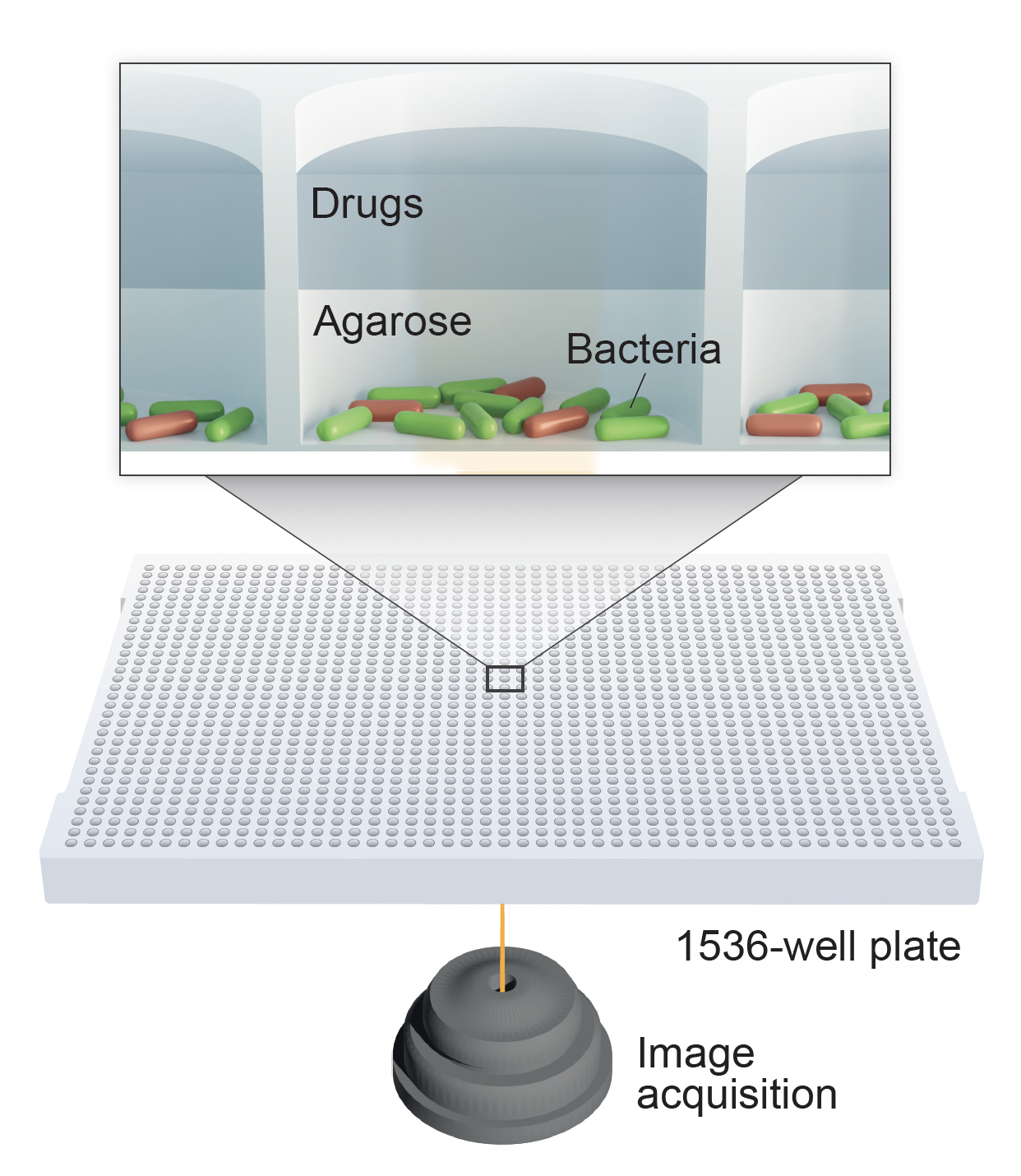
Automation. To systematically study bacterial behaviour and drug responses, we use automated liquid-handling systems that allow us to scale experiments by more than 100-fold. Automation enables us to investigate bacteria at a scale and precision that would be impossible otherwise. By combining automated experimental workflows with custom data analysis pipelines, we generate large-scale datasets to systematically explore how bacteria respond to treatments.
